2. Center for Biomedical Imaging Research, Tsinghua University, Beijing 100084, China
With the progress of science and technology, medicine has entered precision medical era. The phased target of precision medicine is accurate diagnosis, assessment, and treatment of the disease. The rapid development of computer technology promoted the progress of the medical imaging technology and generated the computer assisted surgery. Neuroimaging has played an important role in preoperative evaluation, surgical planning, tumor excision, and functional protection. The three-dimensional (3D) reconstructed images can provide visual information of the spatial relationship of tuomors and their surrounding structures[1,2]. From April of 2015 to December of 2015, this study collected 15 cases of brain tumors and used multimodal image fusion technology to construct brain tumors and their surrounding anatomical structures.
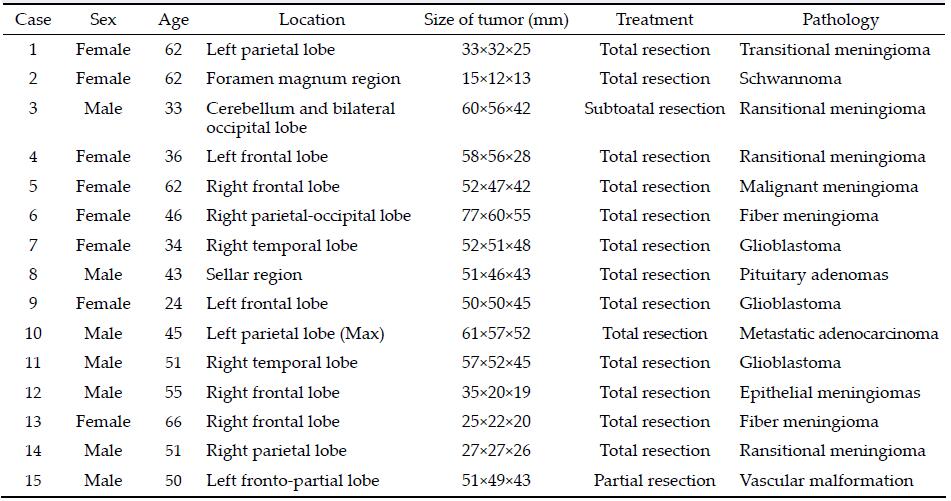
2 Methods 2.1 Basic data of patientsThis study collected 15 cases of brain tumors, 7 cases were male, 8 cases were female, aged 24 to 66 years old. 2 cases of fiber meningioma, 1 case of epithelial meningiomas, 4 cases of transitional meningioma, 1 case of malignant meningioma, 1 case of pituitary adenomas, 3 cases of glioblastoma, 1 case of metastatic tumor, 1 case of vascular malformation, and Schwan-noma in 1 case. Basic data are shown in Table 1.
CT scanning: voltage 120 kV, electric current 319.81 mAs/mA.
MRI scanning: Each patient from Yuquan Hospital of Tsinghua University was underwent collection of MRI scan in biomedial imaging research center. Scan method: (1) MR Scanner: Philips 3.0T Achieva TX; (2)imaging sequence: 3D turbo field echo of inversion pre-pulse excitation (T1WI 3D TFE), 3D T2-weighted inversion recovery based sequence for Cerebrospinal fluid Suppression (CSF) suppression (T2WI 3D FLAIR), 3D phase contrast angiography of vein (3D PCA MRV), bolus tracking of contrast enhance magnetic resonanceangiography (CE-MRA), 3D time of flight magnetic resonance angiography (3D TOF MRA); (3)patients are in a supine position.
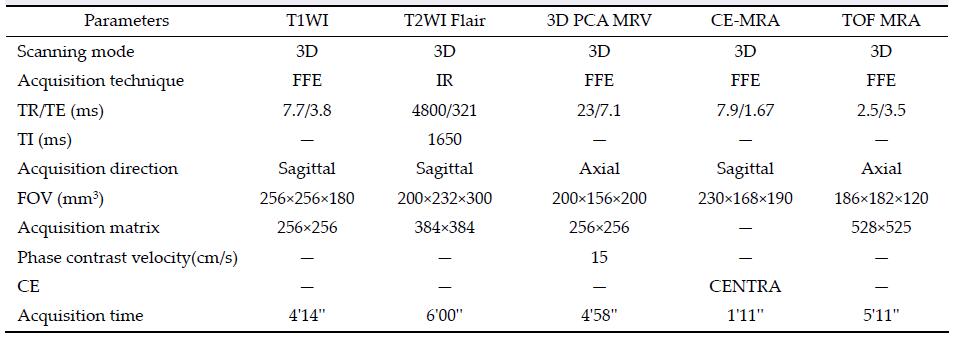
Each patient was underwent collection of CT, T1WI, T2WI Flair, 3D PCA MRV, CE-MRA, TOF MRA, and T1WI enhancement scanning. Specific imaging para-meters are shown in Table 2.
The Dicom image data of 15 patients were processed by NeuroTech software which was developed by Tsinghua University. Data processing contains registra-tion, fusion, and three-dimensional reconstruction. At present, the system has been successfully applied in epilepsy surgery in our hospital. The purpose of this study is to apply this technology to brain tumor surgery.
The registration of MRI scan sequences was adopted by the method of combination of automatic and manual registration. Manual registration used characteristic anatomical symbol (such as eyeball center, the pituitary fossa center and confluence of sinus, etc.) for registra-tion reference point. Each patient was selected with 3-5 reference point for registration.
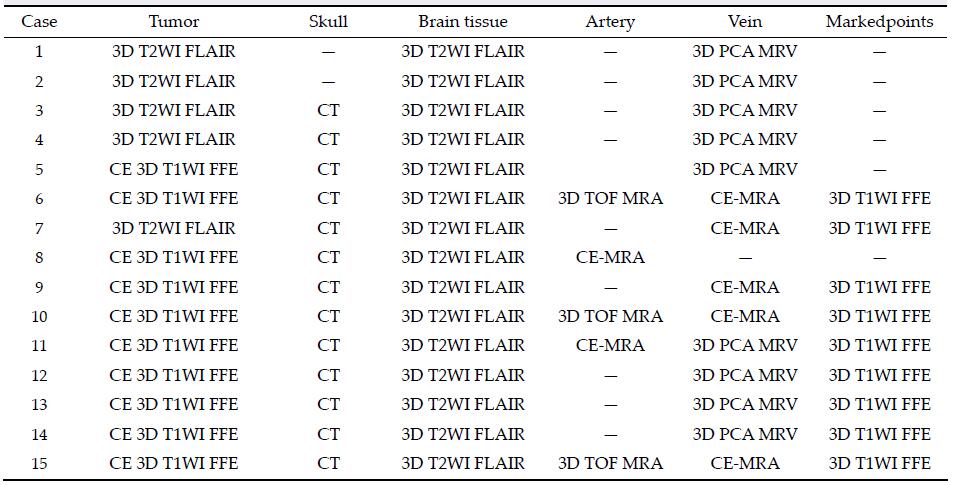
The skull structure has been reconstructed through CT image data and brain tissue by 3D T2WI Flair data, brain tumors by 3D T1WI enhancement scan or
3D T2WI Flair data, marked points by 3D T1WI data. The veins and arteries has been reconstructed through 3D PCA MRV, CE MRA, or 3D TOF MRA data. Specific sources of fusion structures with each patient are shown in Table 3.
Preoperative image reconstruction was obtained in all 15 patients. The reconstructed images such as skull, brain tissue, blood vessels, tumor, and its surrounding strutctures have been successfully reconstructed (Figures 1-8). (1)The skull structure has been recon-structed through CT image data. The 3D reconstructed images can display coronal suture, sagittal suture, and lambdoidal suture clearly. (2)The brain tissue has been reconstructed by 3D T2WI FLAIR. The reconstructed images can clearly display the sulcus, guri, and internal structures. The images also can be cut axially, coronally, or sagittally. (3)The tumor has been reconstructed by 3D T1WI enhancement scan or 3D T2WI Flair data. The reconstructed images clearly show the tumor size, location, shape, and the anatomical relationship of tumor and surrounding structures. (4)The vessels have been reconstructed through 3D PCA MRV, CE MRA, or 3DTOF MRA data, and veins and arteries can be reconstructed clearly and can display the relationship between brain tumor and blood vessels. (5) Marked points have been reconstructed by 3D T1WI FFE.
|
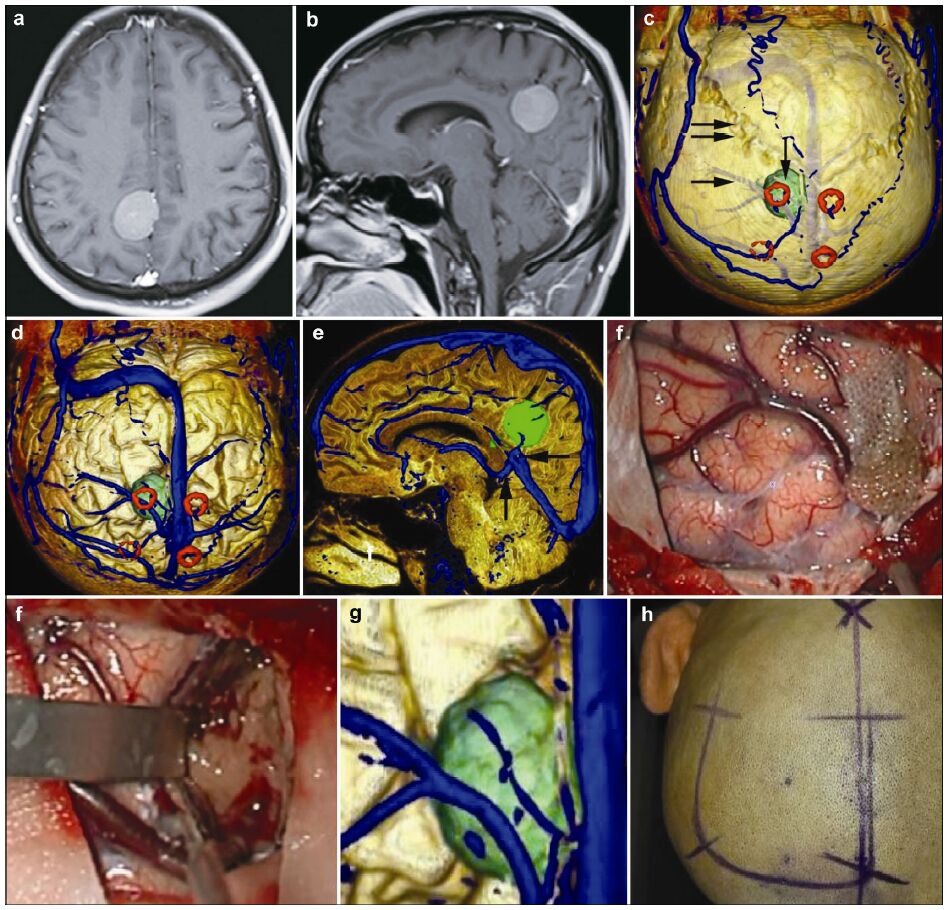
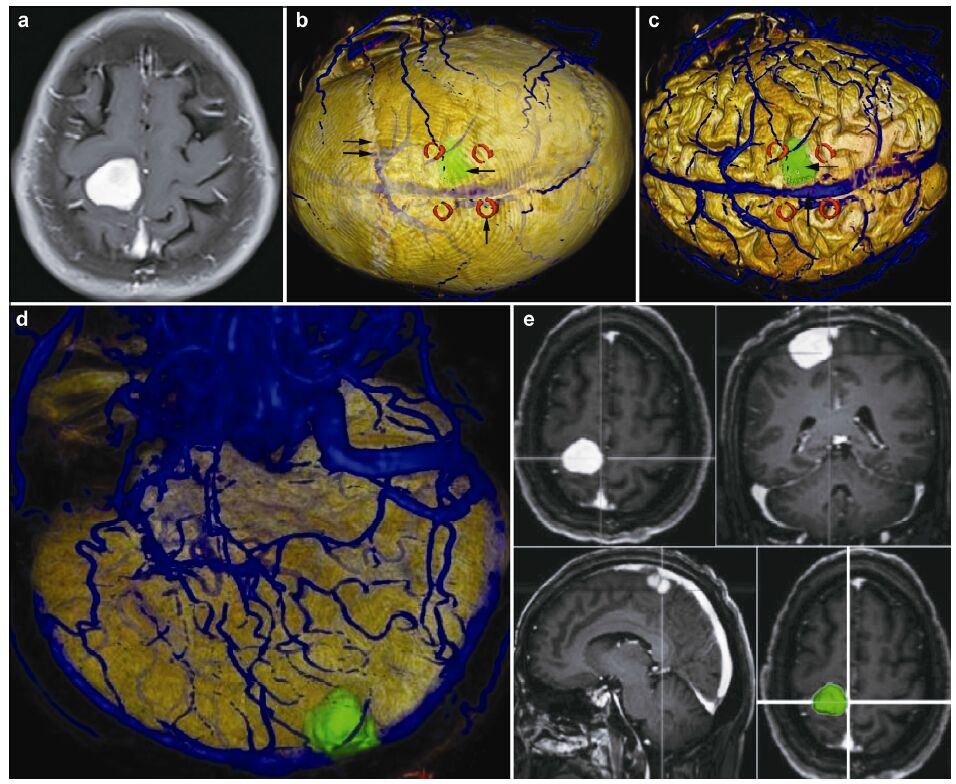
| Figure 1 Magnetic resonance imaging, reconstruction images, surgical incision, and actual operative view in a 51-year-old male patient with a meningioma in right parietal lobe. (a, b) T1 enhancement scans: The tumor presented as homogenous enhancement. (c) 3D reconstruction (skull semitransparent): The tumor (vertical single arrow) located above lambdoidal suture (horizontal double arrow) and the superior cerebral veins (horizontal single arrow) are faintly visible. (d) 3D reconstruction (do not show the skull): On the surface of brain adjacent to the tumor are superior cerebral veins and superior sagittal sinus. (e) Sagittal cutting: The tumor is above the vein of galen and straight sinus. (f, g) The comparison of intraoperative situation and 3D reconstruction: The intraoperative situation is consistent with 3D reconstruction images. (h) Surgical incision. |
|
| Figure 2 Magnetic resonance imaging, reconstruction images, and 3D view of MRI in an 66-year-old female patient with a meningioma in right frontal lobe. (a) T1 enhancement scans: The tumor presented as homogenous enhancement and the precentral gyrus is located anterior to the tumor. (b) 3D reconstruction (skull semitransparent): The tumor (horizontal single arrow) located behind the coronary suture (horizontal double arrow) and the markers (vertical single arrow) are around the tumor. The superior sagittal sinus and superior cerebral veins are faintly visible. (c) 3D reconstruction (do not show the skull): The tumor (horizontal single arrow) is adjacent to the superior sagittal sinus (vertical single arrow). (d) 3D reconstruction (lateral view, brain tissue semitransparent): The tumor is adjacent to the superior sagittal sinus and locates on the surface of the brain. The positional relationship between the tumor and markers can be seen clearly. (e) 3D view: The images of 3D reconstruction are consistent with that of T1 enhancement MRI. |
|
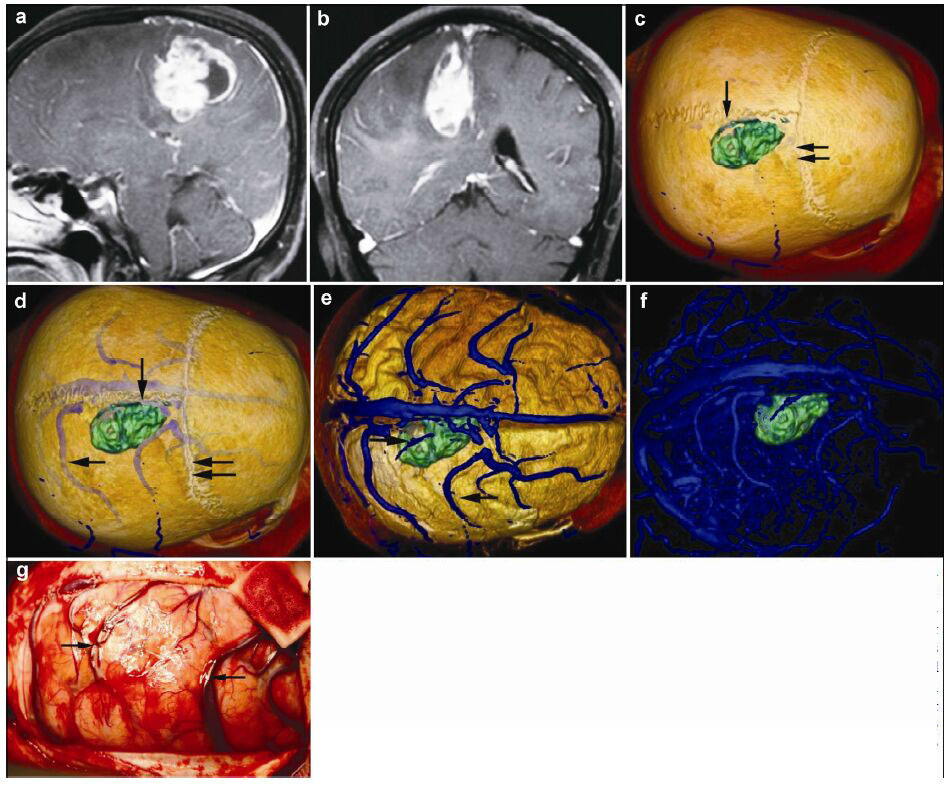
| Figure 3 Magnetic resonance imaging, reconstruction images, and actual operative view in a 62-year-old female patient with a malignant meningioma in right frontal lobe. (a, b) T1 enhancement scans: The tumor presented as heterogeneous enhancement. (c) 3D reconstruc-tion: The tumor located behind the coronary suture (horizontal double arrow) and adjacent to sagittal suture (vertical single arrow). (d)3D reconstruction (skull semitransparent): This picture not only shows the relationship between tumor and sutures (coronary suture, horizontal double arrow; sagittal suture, vertical single arrow) but also shows the relationship between tumor and the superior cerebral veins (horizontal double arrow). (e) 3D reconstruction (do not show the skull): The tumor locates between two superior cerebral veins (horizontal single arrow). (f) 3D reconstruction: This picture only shows the tumor and veins. (g) Intraoperative situation: The intraoperative situation is consistent with 3D reconstruction images. |
|
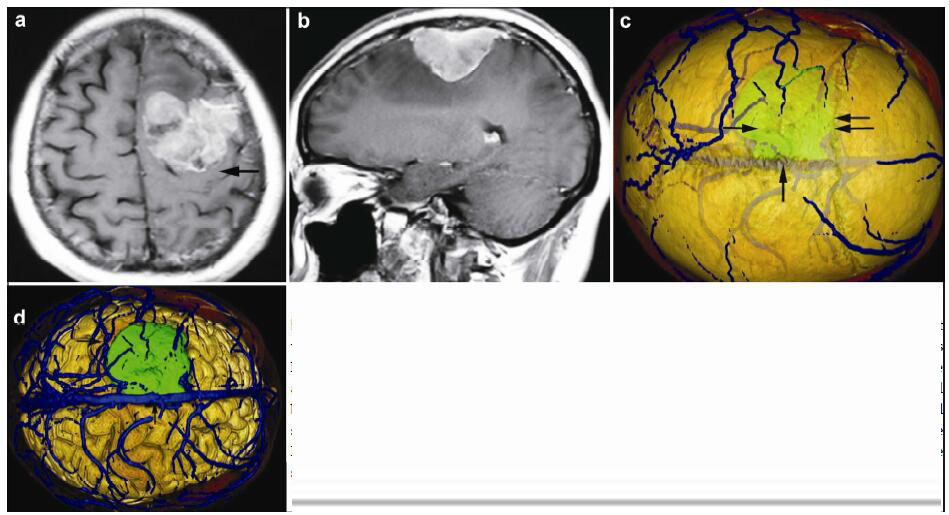
| Figure 4 Magnetic resonance imaging and reconstruction images in a 36-year-old female patient with a meningioma in left frontal lober. (a, b) T1 enhancement scans: The tumor presented as heterogeneous enhancement and locates anerior to left postcentral gyrus (horizontal single arrow). (c) 3D reconstruction (skull semitransparent): The tumor (horizontal single arrow) located behind the coronary suture (horizontal double arrow) and adjacent to sagittal suture (vertical single arrow). (d) 3D reconstruction (do not show the skull): The tumor locates anterior to the left postcentral gyrus (vertical single arrow) and the right side of the tumor is adjacent to the superior sagittal sinus (vertical single arrow). |
|
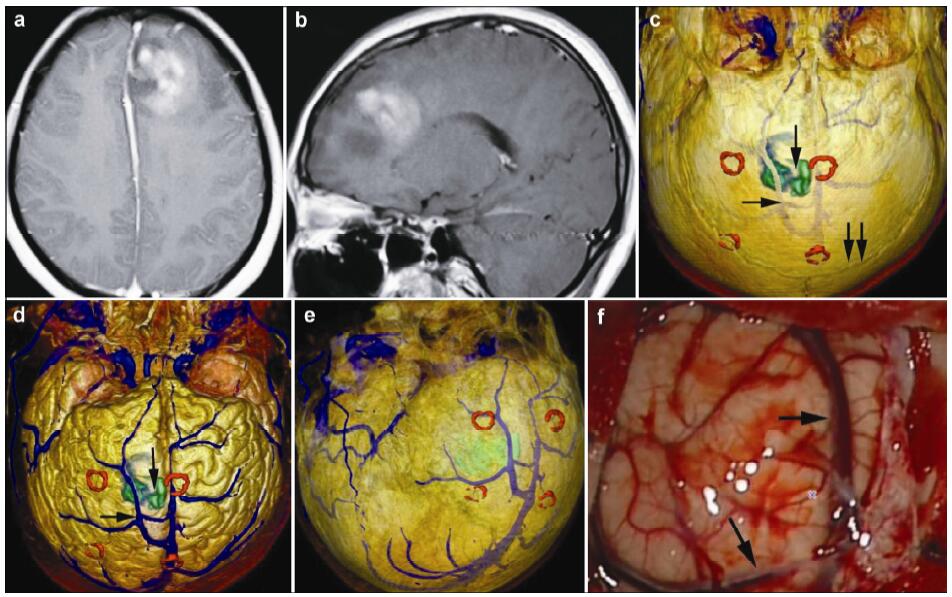
| Figure 5 Magnetic resonance imaging, reconstruction images, and actual operative view in a 24-year-old female patient with a glioblastoma in left frontal lobe. (a, b) T1 enhancement scans: The tumor presented as heterogeneous enhancement. (c) 3D reconstruction: 3D reconstruction (skull semitransparent): This picture not only shows the relatioshop between tumor and sutures (vertical double arrow) but also shows the relatioshop between tumor (vertical single arrow) and the superior cerebral veins (horizontal single arrow). (d) 3D reconstruction (do not show the skull): The superior cerebral veins are shown clearly and the tumor (vertical single arrow) is under one large superior cerebral vein (horizontal single arrow). (e) 3D reconstruction (superolateral view, skull, and brian tissue semitransparent): In this picture, we can see tumor, markers, superior cerebral veins, and brain tissue simultaneously. (f) Intraoperative situation: The intraoperative situation is consistent with 3D reconstruction images. |
|
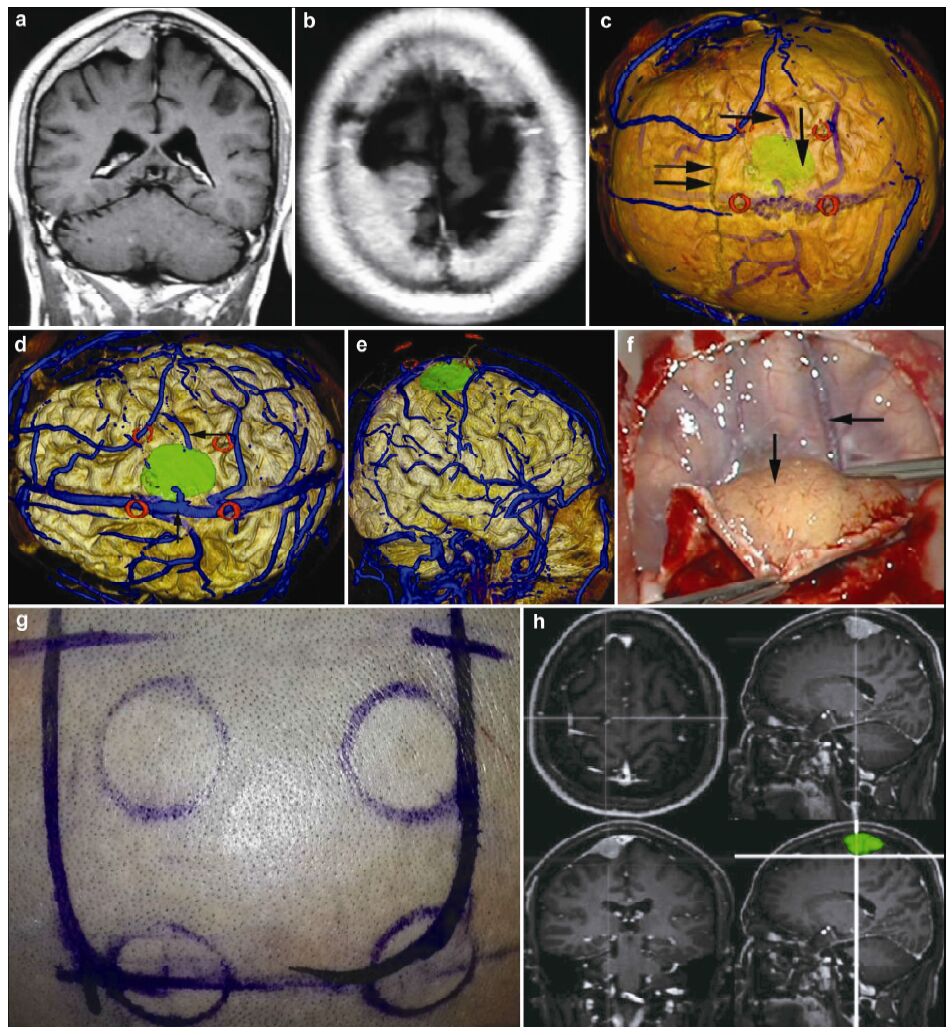
| Figure 6 Magnetic resonance imaging, reconstruction images, surgical incision, 3D view, and actual operative view in a 55-year-old male patient with a meningioma in right frontal lobe. (a, b) T1 enhancement scans: The tumor presented as homogenous enhancement. (c) 3D reconstruction (skull semitransparent): The tumor (vertical single arrow) located behind the coronary suture (horizontal double arrow) and adjacent to sagittal suture. The right central sulcus vein (horizontal single arrow) is under the tumor. (d) 3D reconstruction (do not show the skull): The tumor is adjacent to the superior sagittal sinus (vertical single arrow) and superior sagittal sinus (vertical single arrow). (e) 3D reconstruction (lateral view): The tumor is adjacent to the superior sagittal sinus and locates on the surface of the brain above the right central sulcus vein (horizontal single arrow). The markers are around the tumor. (f) Intraoperative situation: The intraoperative situation is consistent with 3D reconstruction images. (g) Surgical incision. (h) 3D view: The images of 3D reconstruction are consistent with that of T1 enhancement MRI. |
|
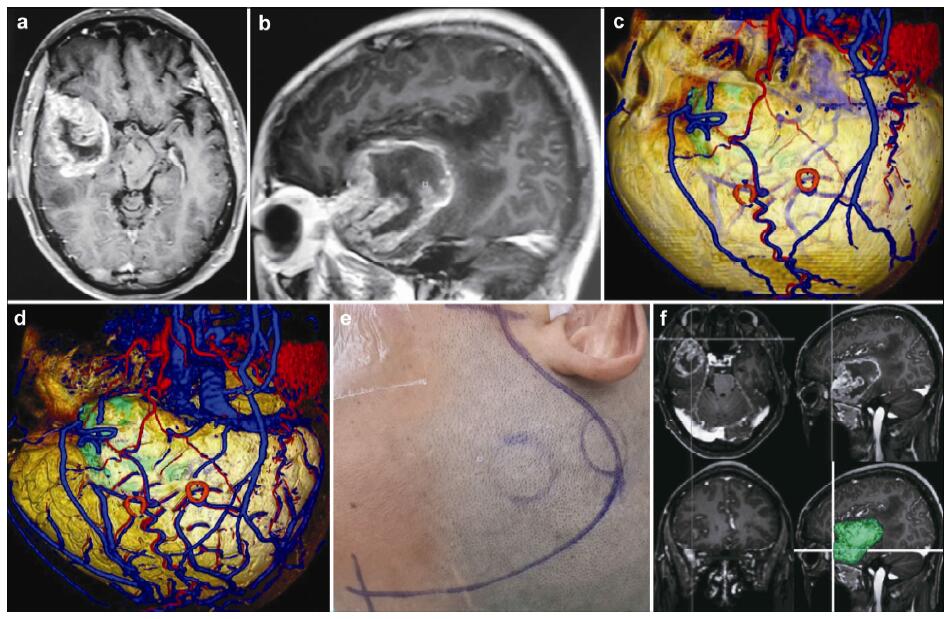
| Figure 7 Magnetic resonance imaging, reconstruction images, surgical incision, and 3D view in a 51-year-old male patient with a glioblastoma in right frontal lobe. (a, b) T1 enhancement scans: The tumor presented as heterogeneous enhancement. (c) 3D reconstruction (skull semitransparent): The scalp blood vessels have been shown clearly. The tumor and superior cerebral veins are faintly visible. (d) 3D reconstruction (lateral view, do not show the skull): The tumor located under the markers. The veins, arteris, brian surface, and markers have shown clearly and the tumor is faintly visible. (e) Surgical incision. (f) 3D views: The images of 3D reconstruction are consistent with that of T1 enhancement MRI. |
|
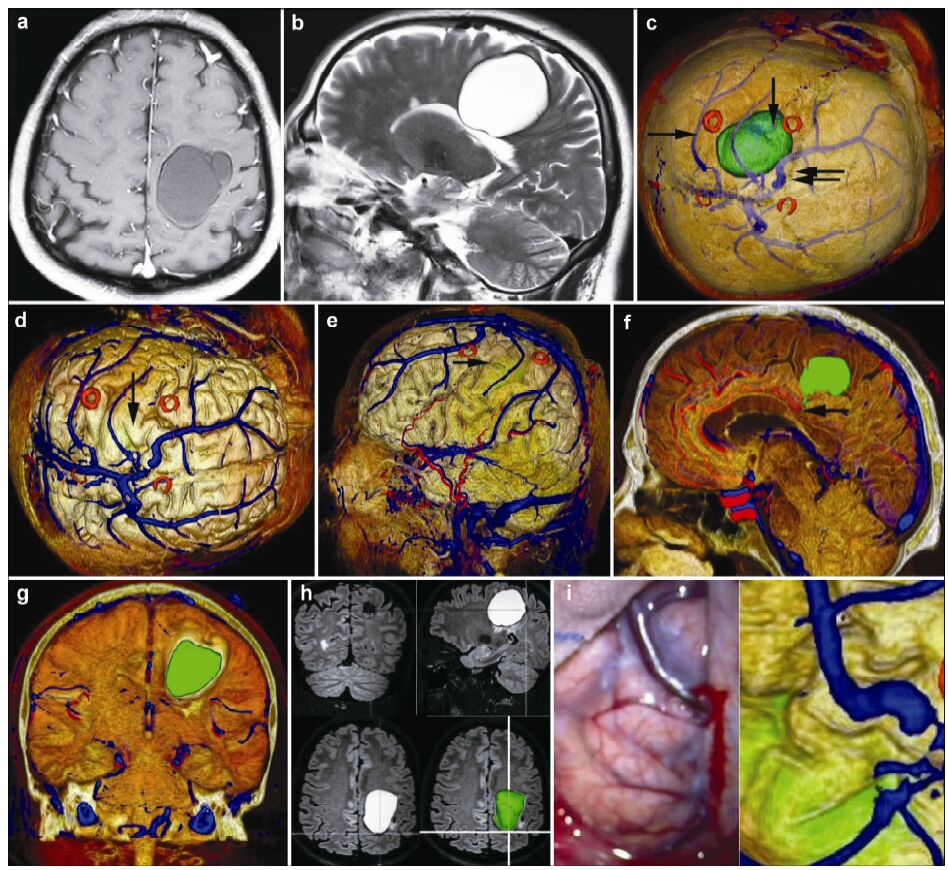
| Figure 8 Magnetic resonance imaging, reconstruction images, surgical incision, 3D view, and actual operative view in a 51-year-old male patient with a vascular malformation in right frontal lobe. (a, b) T1 enhancement scan: The cyst presented as no enhancement; (b) T2 scan: The cyst presented as high signal. (c) 3D reconstruction (skull semitransparent): The cyst (vertical single arrow) located behind the coronary suture (horizontal double arrow) and the markers are around the cyst. The superior sagittal sinus and superior cerebral veins (horizontal single arrow) are faintly visible. (d) 3D reconstruction (do not show the skull): The cyst (vertical single arrow) is faintly visible and locates anterior to one superior cerebral vein. The veins on the surface of the brain can be seen clearly. (e) 3D reconstruction (lateral view): The cyst (horizontal single arrow) is anterior to one superior cerebral vein and locates under the motor cortex. (f) Sagittal cutting: The corpus callosum (horizontal single arrow) is compressed by the cyst. (g) Coronay cutting: The lateral ventricle and corpus callosum are compressed by the cyst. (h) 3D view: The images of 3D reconstruction are consistent with that of flair MRI. (i) The comparison of intraoperative situation and 3D reconstruction: The intraoperative situation is consistent with 3D reconstruction images. |
The reconstructed images are consistent with MRI images and the real anatomical morphology in the process of operation. The whole 3D structures including tumor, brain surface, skull, and vessels were success-fully reconstructed based on CT and MRI images. Reconstruction of the structures can be any combina-tion, adjust color, through the rotation images can be observed from any angle. Reconstruction of brain tissue and skull can be semi transparent to display deep structures. Reconstruction of the structures can be axial, coronal, and sagittal cutting to show relationship among tumor and surrounding structures. The recon-structed 3D structures clearly depicted the tumor features, such as size, location, and shape, and provided a visual information of the spatial relationship among its surrounding structures. The images of deep tumors can be projected onto skull surface, and show the relationship between tumor and skull anatomic landmarks such as coronal suture, lambdoid suture, or sagittal suture.
We can hide any reconstructed images such as skull, brain tissue, blood vessles, or tumors to facilitate the display of interested areas or structures. Based on 3D reconstructedimages, we can design incision and surgical approach more accurately, suitably, and individually.
3.2 Postoperative image reconstructionThe postoperative reconstructed images have been obtained in 3 patients. The reconstructed images such as skull, brain tissue, blood vessels, and its surrounding structures have been reconstructed clearly. (1)The 3D images of brain tissue have been constructed through 3D T2WI FLAIR data. Cortical structures of operation area have not been displayed clearly due to edema. (2)Vascular reconstruction: The veins have been recon-structed by 3D PCA MRV data. The postoperative constructed images clearly display that the central veins have been in good protection. The reconstructed images provided local anatomic relationship between vessels and other structures. Vessels were clearly displayed in 3D. So the vessels were more likely to be protected.
3.3 Surgical results and complications(1) Tumor resection degree: 13 patients were underwent total excision with microscope, 1 case of subtotal resection, 1 case of partial resection. All patients showed no residual lesions except 1 case with partial residual tumor and 1 case with cyst obviously narrowed. (2)Complications: Seven patients with postoperative recovery of myodynamia, 1 patient with recovery of paresthesia, 2 patients with language function im-proved, 2 patients with visual acuity improved, and 1patient developed into deep coma, 1 patient with epileptic seizure.
3.4 Follow-up resultsFollow-up time was 2 months to 1 year. Regular MRI examination showed no tumor recurrence except one malignant meningioma. 1 patient died after opera-tion in one month. The rest of the patients didn’t developed new symptoms.
4 DiscussionBrain tumor and its surrounding structures are relati-vely complex. Both CT and MRI examination are single-modal image technology which cannot meet the need of clinical practice. We urgently want a new technology which can display brain tumor and its surrounding anatomical structures clearly. Multi- modal image technology is a computer-generated 3D technology that provides interactivity for the user[3]. The principles of CT and MRI imaging are different and CT and MRI images can provide different information[4]. Different imaging methods are com-plementary and provide more information than each imaging method. Against this background, medical applications of multi-modal image technology have been developing rapidly, ranging from a research to clinical application. With the help of those technologies, the neurosurgeons are able to process large amounts of image data and to create accurate and realistic 3D reconstructions of anatomic structures and related tumors. So we can obtain 3D reconstructed images which neurosurgeons can explore the brain tumors for guiding the design of incision, choosing the operative approaches, and resecting the tumors. The 3D recon-structed images allowed neurosurgeons to facilitate and increase understanding of the complex anatomic and pathological relationships of the tumor. The pre-operative information provided by 3D reconstructed images was useful and helpful during the operative procedure[2-8].
The multimodal image technology has been applied in various fields of neurosurgery[5-13]. The reconstructed 3D images provided the relationship among tumor, vascular, skull, and marked points, and allowed neuro-surgeons to facilitate the incision design, the choice of surgical approach, and avoid important nerve and vascular injury. This technology has an important supporting role in operative procedure. Saito et al.[14] reported that surgical simulations of cerebrovascular disease with multimodal fusion 3D images are very helpful. They utilize 3D images to help clipping of aneurysm. According to the fusion images, they can simulate different surgical approaches and select the best one. Mert[14] displayed the relationship among brain tumor, superior cerebral veins, brain surface, the foramen of monro, anterior vein of septum pellucidum, striothalamate vein, choroid plexus, and septum pellucidum according to the multi-modal image fusion technology. They used those fusion images to design the surgical approach and resect the tumor in order to remove tumor more accurately and damage the brain tissue more slightly.
In this research case 5 (Figure 3), the 3D recon-structed images can clearly shows that the tumor Figure 3 located posterior to the coronal suture and medial to the sagittal suture. The tumor also located between two central veins at the same time, and we can select this space to remove the tumor in order to avoid the damage of important veins. In case 14 (Figure 1), the 3D reconstructed images display that the tumor near the confluence of bilateral lambdoid sutures. Below the tumor are straight sinus and great cerebral veins, and there is a vein in the surface of brain tissue posterior the tumor. In this situation, when we design the incision and bone flap we must avoid this vein in order to remove the tumor more easily. So we don’t design theincision which is to be nearest to the tumor. We select the approach between the vein and superior sagittal sinus. When we open the flap, see the confluence of bilateral lambdoid sutures, we know where the tumor and vein are. We want to look through the surface and see where various structures are located. In this situation, we have developed a see-throught-X-ray vison. This may minimize surgical damage and avoid vein injury. In case 9 (Figure 5), the 3D reconstructed images shows that there is a vein on the surface of the brain above the tumor. When we design the bone flap, we can avoid the flap right above the tumor. We design the flap lateral to the vein, so we can protect this vein after the removal of the tumor.
In case 10, the first third of the sagittal sinus is absent, and it has been replaced by massive venous drainage on both sides. In this case, we should avoid to damage those veins. When we incise the dura mater, we incise the dura mater from lateral to medial, because the terminal superior cerebral veins are closely related to dure mater. When we incise small part of dura mater we ensure that there are no big venous drainage. Then we gradually open the dura mater in order to avoid damaging important veins. In some cases, the lateral areas also have big veins, which we often cann’t easily foresee. By contrast, 3D recon-structed images can provide us the whole course of veins. 3D reconstructed images also can clearly show the normal and mutant brain veins. Before we incise the mater, we clearly knew where those veins are. In case 12 (Figure 6), 3D reconstructed images clearly show that one central vein is first located under the tumor and then flows into the superior sagittal sinus medial to the tuomor. Traditional 2D images cann’t clearly display the whole course of this vein. 3D reconstructed images clearly show the course of the vein and its relationship with the tumor and superior sagittal sinus. During removal of the tumor, we fairly know the anatomic relation of the tumor, vein, and gyri. In many cases, there are abnormal venous drainage near the tumor region. With help of fusion images, we can reduce the possibility of vein injury. In case 8, the 3D reconstructed images clearly show the relationship among the tumor, anterior cerebral artery, and anterior communicating artery. The tumor embraced the anterior communicating artery and part of A1, A2 segment of anterior cerebral artery. When we perform an operation to remove the tumor, we clearly know the anatomic relation of these structures. So we can reduce the probability of injury of those arteries, otherwise can lead to serious complications.
In a lot of low grade glioma, the brain surface struc-ture is normal, so the intraoperative localization is difficult. Mert[14] has used image fusion technology to guide the incision low grade gliomas. They accurately positioned tumor according to the 3D reconstructed images which show the relationship between tumor and its surrounding blood vessels and the cortex. With aid of this technology, they reduced the neural electrophysiological detection region and save the time of surgery. In this study case 7, the tumor located temporal lobe and the brain surface is relatively nor-mal. The 3D reconstructed images clearly show the relationship between tumor and veins. The tumor located behind the trolard vein and anterior to the labbe veins. The images also show the relationship between tumor and other small veins. So we can precisely predict the location of the tumor. During removal of the tumor, we can reduce the normal cortex damage and achieve the precise individualized tumor resection.
Multimodal image fusion technology has a good application prospect but there are some problems. NeuroTech software is used in this study for data fusion. The 3D reconstructed images of the brain tissue, blood vessels, bones, and tumors have been relatively clear, but it is impossible to fuse DTI data. Secondly, in terms of the reconstruction of the blood vessels, we use a variety of data sources such as 3D PCA MRV, CE MRA, or TOF MRA to reconstruct the vessels but there is no one method which can clearly display more veins and arteries respectively. Third, for the reconstruction of the tumor, we often use 3D T1WI to reconstruct the images. When the tumor enhancement was more obvious, such as meningioma, reconstruction result is better. However, when the tumor enhancement was slight or no enhancement, the reconstruction of tumor was difficult. Fourth, when we reconstruct the tumor, some vessels which related the tumor can also be reconstructed. So the tumor reconstruction is not very exact.
Conflict of interests
All contributing authors have no conflict of interests.
| [1] | da Rocha LG, Amaro JE. Seeking tools for image fusion between computed tomography, structural and functional magnetic resonance methods for applications in neurosurgery. Einstein (São Paulo) , 2012, 10 (2) : 151–157. DOI:10.1590/S1679-45082012000200007 |
| [2] | Kin T, Nakatomi H, Shojima M, Tanaka M, Ino K, Mori H, Kunimatsu A, Oyama H, Saito N. A new strategic neurosurgical planning tool for brainstem cavernous malformations using interactive computer graphics with multimodal fusion images. J Neurosurg , 2012, 117 (1) : 78–88. DOI:10.3171/2012.3.JNS111541 |
| [3] | Oishi M, Fukuda M, Yajima N, Yoshida K, Takahashi M, Hiraishi T, Takao T, Saito A, Fujii Y. Interactive presurgical simulation applying advanced 3D imaging and modeling techniques for skull base and deep tumors. J Neurosurg , 2013, 119 (1) : 94–105. DOI:10.3171/2013.3.JNS121109 |
| [4] | Saito N, Kin T, Oyama H, Yoshino M, Nakagawa D, Shojima M, Imai H, Nakatomi H. Surgical simulation of cerebrovascular disease with multimodal fusion 3-dimensional computer graphics. Neurosurgery , 2013, 60 (Suppl 1) : 24–29. |
| [5] | Inoue HK, Nakajima A, Sato H, Noda SE, Saitoh J, Suzuki Y. Image fusion for radiosurgery, neurosurgery and hypofractionated radiotherapy. Cureus , 2015, 7 (3) : e252. |
| [6] | Ferroli P, Tringali G, Acerbi F, Schiariti M, Broggi M, Aquino D, Broggi G. Advanced 3-dimensional planning in neurosurgery. Neurosurgery , 2013, 72 (Suppl 1) : 54–62. |
| [7] | Kockro RA, Reisch R, Serra L, Goh LC, Lee E, Stadie AT. Image-guided neurosurgery with 3-dimensional multimodal imaging data on a stereoscopic monitor. Neurosurgery , 2013, 72 (Suppl 1) : 78–88. |
| [8] | Cardinale F, Pero G, Quilici L, Piano M, Colombo P, Moscato A, Castana L, Casaceli G, Fuschillo D, Gennari L, et al. Cerebral angiography for multimodal surgical planning in epilepsy surgery: Description of a new three-dimensional technique and literature review. World Neurosurg , 2015, 84 (2) : 358–367. DOI:10.1016/j.wneu.2015.03.028 |
| [9] | Liu WB, Guo H, Du XF, Zhou WJ, Zhang GM, Ding H, Wang GZ. Cortical vessel imaging and visualization for image guided depth electrode insertion. Comput Med Imaging Graph , 2013, 37 (2) : 123–130. DOI:10.1016/j.compmedimag.2012.05.004 |
| [10] | Takai K, Kin T, Oyama H, Iijima A, Shojima M, Nishido H, Saito N. The use of 3D computer graphics in the diagnosis and treatment of spinal vascular malformations. J Neurosurg Spine , 2011, 15 (6) : 654–659. DOI:10.3171/2011.8.SPINE11155 |
| [11] | Kin T, Shin M, Oyama H, Kamada K, Kunimatsu A, Momose T, Saito N. Impact of multiorgan fusion imaging and interactive 3-dimensional visualization for intraventricular neuroendoscopic surgery. Neurosurgery , 2011, 69 (1 Suppl Operative) : ons40–48. |
| [12] | Kin T, Oyama H, Kamada K, Aoki S, Ohtomo K, Saito N. Prediction of surgical view of neurovascular decompression using interactive computer graphics. Neurosurgery , 2009, 65 (1) : 121–129. DOI:10.1227/01.NEU.0000347890.19718.0A |
| [13] | Korvenoja A, Kirveskari E, Aronen HJ, Avikainen S, Brander A, Huttunen J, Ilmoniemi RJ, Jääskeläinen JE, Kovala T, Mäkelä JP, et al. Sensorimotor cortex localization: Comparison of magnetoencephalography, functional MR imaging, and intraoperative cortical mapping. Radiology , 2006, 241 (1) : 213–222. DOI:10.1148/radiol.2411050796 |
| [14] | Mert A, Buehler K, Sutherland GR, Tomanek B, Widhalm G, Kasprian G, Knosp E, Wolfsberger S. Brain tumor surgery with 3-dimensional surface navigation. Neurosurgery , 2012, 71 (2 Suppl Operative) : 286–295. |